
Rice Science ›› 2018, Vol. 25 ›› Issue (6): 297-307.DOI: 10.1016/j.rsci.2018.06.008
• Research Papers • Next Articles
Ran Qin1, Akhter Delara1,2, Chengcong Yang1, Kumar Nath Ujjal3, Eshag Jamal1, Xiaoli Jin1, Chunhai Shi1( )
)
Received:2018-04-02
Accepted:2018-06-14
Online:2018-11-28
Published:2018-08-20
Ran Qin, Akhter Delara, Chengcong Yang, Kumar Nath Ujjal, Eshag Jamal, Xiaoli Jin, Chunhai Shi. SRG1, Encoding a Kinesin-4 Protein, Is an Important Factor for Determining Grain Shape in Rice[J]. Rice Science, 2018, 25(6): 297-307.
Add to citation manager EndNote|Ris|BibTeX
| Trait | Wild type | srg1 mutant |
|---|---|---|
| Grain length (mm) | 7.51 ± 0.37 | 6.08 ± 0.29** |
| Grain width (mm) | 3.24 ± 0.14 | 3.53 ± 0.18* |
| Grain length/width | 2.32 ± 0.17 | 1.72 ± 0.13** |
| Roundness value | 0.46 ± 0.12 | 0.61 ± 0.21** |
| No. of grains per panicle | 204.4 ± 10.5 | 114.0 ± 12.0** |
| 1000-grain weight (g) | 25.35 ± 0.86 | 21.44 ± 0.63* |
| Plant height (cm) | 83.90 ± 1.45 | 50.35 ± 1.22** |
| Seed-setting rate (%) | 92.6 ± 1.1 | 87.1 ± 0.4* |
| Panicle length (cm) | 20.21 ± 0.98 | 13.29 ± 0.44** |
Table 1 Agronomic traits of wild type and srg1 mutant.
| Trait | Wild type | srg1 mutant |
|---|---|---|
| Grain length (mm) | 7.51 ± 0.37 | 6.08 ± 0.29** |
| Grain width (mm) | 3.24 ± 0.14 | 3.53 ± 0.18* |
| Grain length/width | 2.32 ± 0.17 | 1.72 ± 0.13** |
| Roundness value | 0.46 ± 0.12 | 0.61 ± 0.21** |
| No. of grains per panicle | 204.4 ± 10.5 | 114.0 ± 12.0** |
| 1000-grain weight (g) | 25.35 ± 0.86 | 21.44 ± 0.63* |
| Plant height (cm) | 83.90 ± 1.45 | 50.35 ± 1.22** |
| Seed-setting rate (%) | 92.6 ± 1.1 | 87.1 ± 0.4* |
| Panicle length (cm) | 20.21 ± 0.98 | 13.29 ± 0.44** |
| ID | Gene function annotation |
|---|---|
| LOC_Os09g02650 | Kinesin motor domain containing protein, putative, expressed |
| LOC_Os09g02670 | Expressed protein |
| LOC_Os09g02680 | Retrotransposon protein, putative, unclassified |
| LOC_Os09g02690 | Expressed protein |
| LOC_Os09g02700 | Polyadenylate-binding protein, putative, expressed |
| LOC_Os09g02710 | CBS domain containing membrane protein, putative, expressed |
| LOC_Os09g02729 | Phospholipase C, putative, expressed |
| LOC_Os09g02750 | Expressed protein |
| LOC_Os09g02760 | Retrotransposon protein, putative, Ty1-copia subclass, expressed |
| LOC_Os09g02770 | Translation initiation factor IF-2, putative, expressed |
| LOC_Os09g02780 | OsMADS77-MADS-box family gene with M-alpha type-box, expressed |
Table 2 Functional predications of candidate genes of srg1.
| ID | Gene function annotation |
|---|---|
| LOC_Os09g02650 | Kinesin motor domain containing protein, putative, expressed |
| LOC_Os09g02670 | Expressed protein |
| LOC_Os09g02680 | Retrotransposon protein, putative, unclassified |
| LOC_Os09g02690 | Expressed protein |
| LOC_Os09g02700 | Polyadenylate-binding protein, putative, expressed |
| LOC_Os09g02710 | CBS domain containing membrane protein, putative, expressed |
| LOC_Os09g02729 | Phospholipase C, putative, expressed |
| LOC_Os09g02750 | Expressed protein |
| LOC_Os09g02760 | Retrotransposon protein, putative, Ty1-copia subclass, expressed |
| LOC_Os09g02770 | Translation initiation factor IF-2, putative, expressed |
| LOC_Os09g02780 | OsMADS77-MADS-box family gene with M-alpha type-box, expressed |
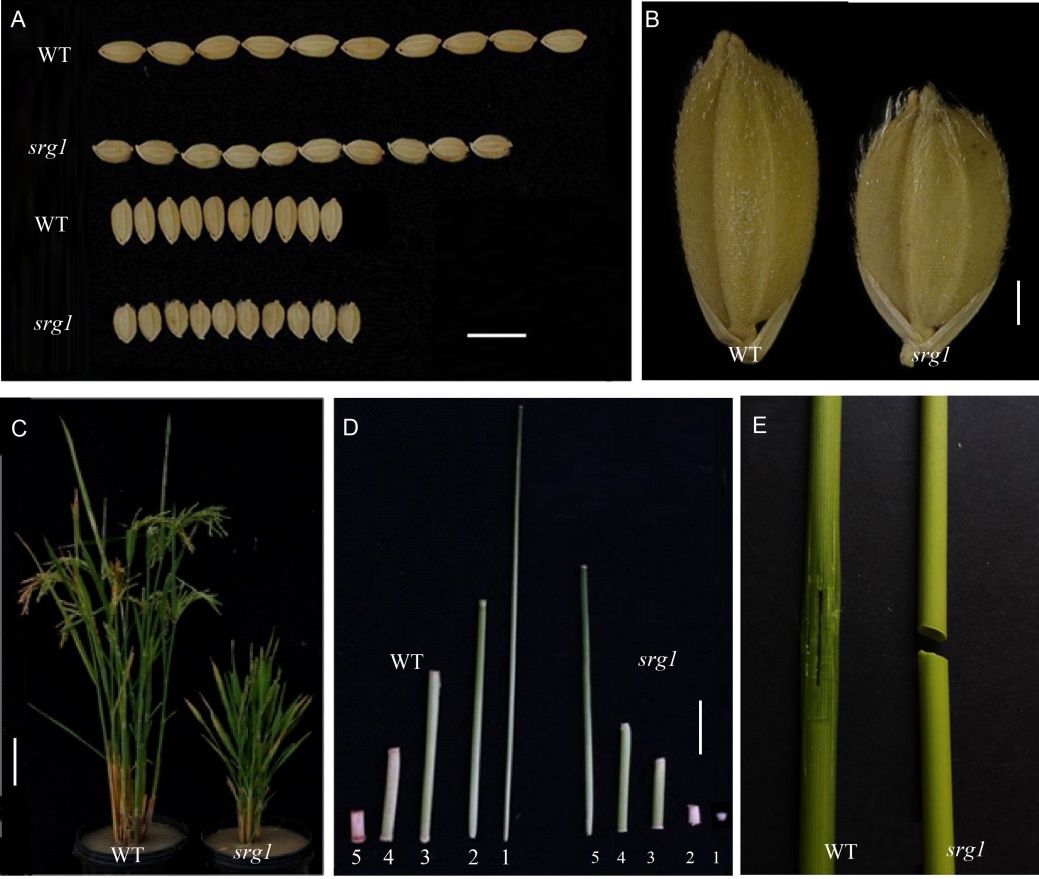
Fig. 1. Phenotypes of wild type (WT) and srg1 mutant. A, Grain length. B, Grain width. C, Plant height. D, Internode. The numbers 1, 2, 3, 4 and 5 refer to the 1st, 2nd, 3rd, 4th and 5th internodes from the top, respectively. E, Culm brittleness. Scale bars in A, B, C and D are 1.0 cm, 0.1 mm, 10.0 cm and 2.5 cm, respectively.
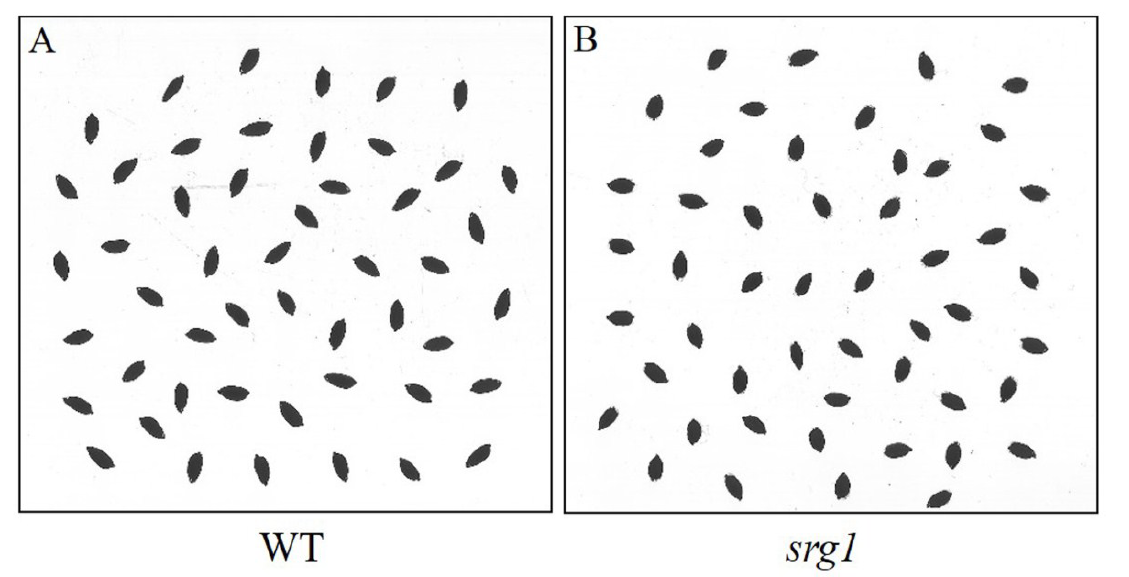

Supplemental Fig. 1. The scanned images of the grain shape of WT(A) and srg1(B). Using the images the roundness was obtained by the rice appearance quality inspection system(SC-E).
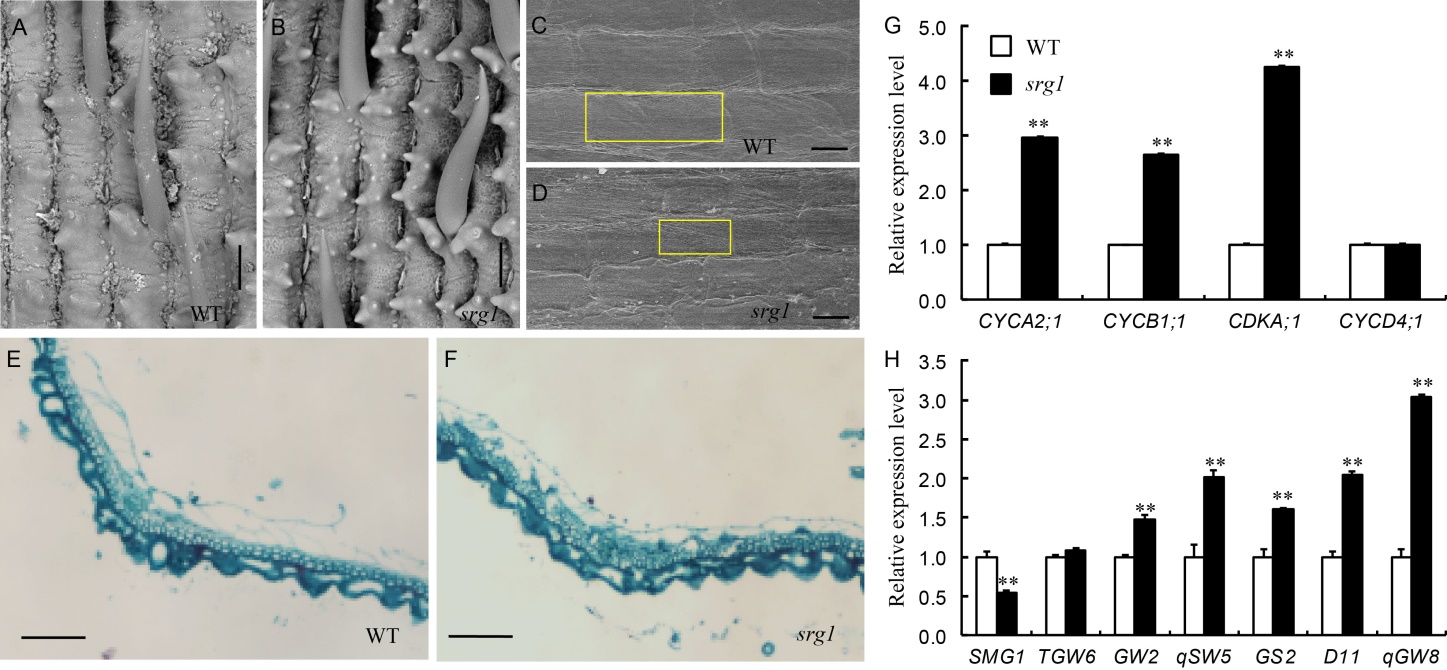
Fig. 2. Effect of SRG1 on grain shape. A-D, Scanning electron microscope analysis of outer and inner surfaces of wild type (WT, A and C) and srg1 mutant (B and D) lemmas, respectively. Square frame in C and D indicates the cell length. E and F, Cross-sections of the central parts of WT and srg1 lemmas. G, Expression levels of cell cycle genes in WT and srg1 spikelets from young panicles. H, Expression levels of genes related to grain shape in WT and srg1 spikelets from young panicles. Rice Actin gene was used as an internal control. Scale bars in A, B, C, D, E and F are 50 µm, 50 µm, 25 µm, 25µm, 100 µm and 100 µm, respectively. Values represent mean ± SD (n = 3); **, P ≤ 0.01.
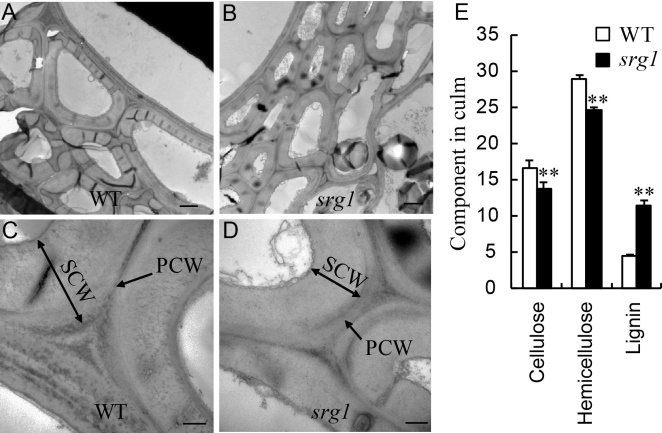
Fig. 3. Alterations of structure and components in cell wall of culm in wild type (WT) and srg1 at heading stage. A-D, Transverse electric and magnetic observation of sclerenchyma cell walls in wild type (WT) (A and C) and srg1 culms (B and D). C and D were the amplification of cell wall structure in A and B, respectively. E, Cell wall main components in culm of WT and srg1. PCW, Primary cell wall; SCW, Secondary cell wall.Scale bars in A, B, C and D are 2.0 µm, 2.0 µm, 0.5 µm and 0.5 µm, respectively. Value represents the mean ± SD (n = 3); **, P ≤ 0.01.
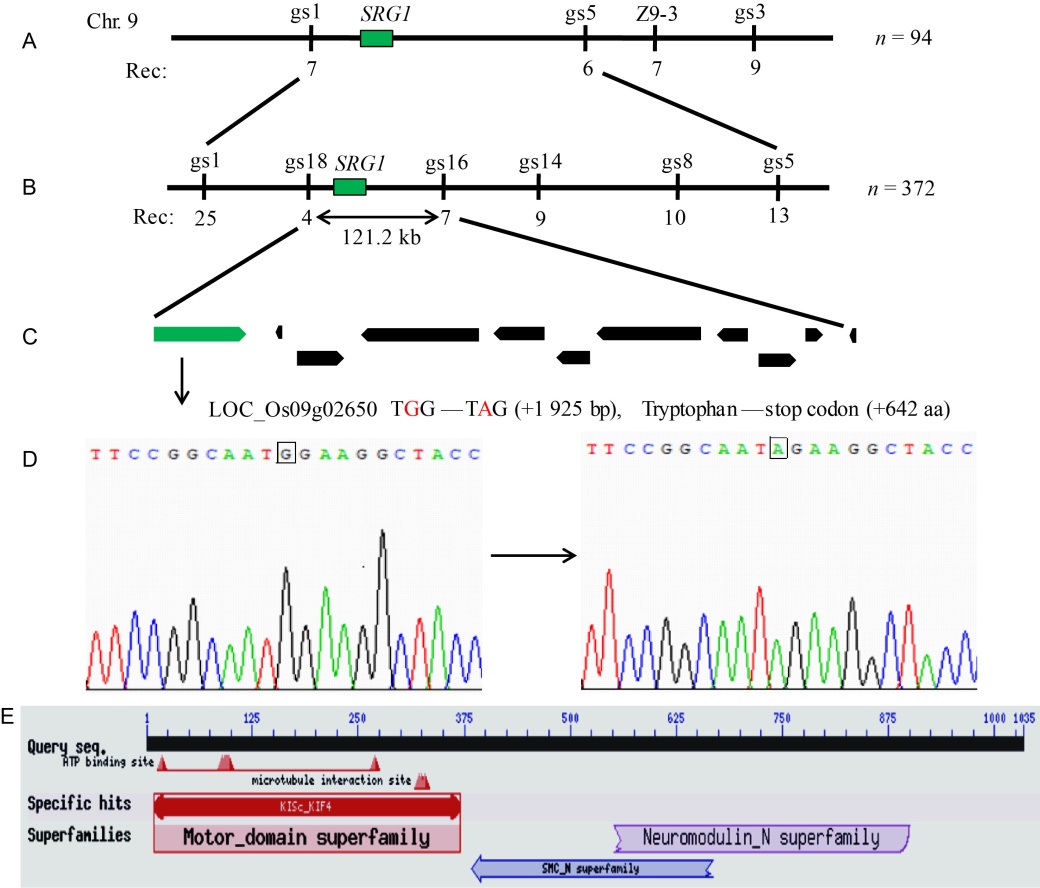
Fig. 4. Map-based cloning of SRG1.A, Primary mapping of srg1 using 94 F2 individuals. The recombinant number was listed under the corresponding marker. B, SRG1 was fine-mapped to a 121.2 kb interval between markers gs18 and gs16 with 372 F2 individuals. C, Total 11 genes were annotated in the region according to the database of Rice Genome Annotation Project. LOC_Os09g02650 with the green color was presumed to be the locus of SRG1. D, The sequencing analysis of LOC_Os09g02650. There was 1 bp substitution (G to A) occurred in the 16th exon (+1 925 bp) of the srg1, which caused the tryptophan (+642 aa, TGG) changing to a stop codon (TAG). E, The structure domains of SRG1 obtained from the National Center for Biotechnology Information.
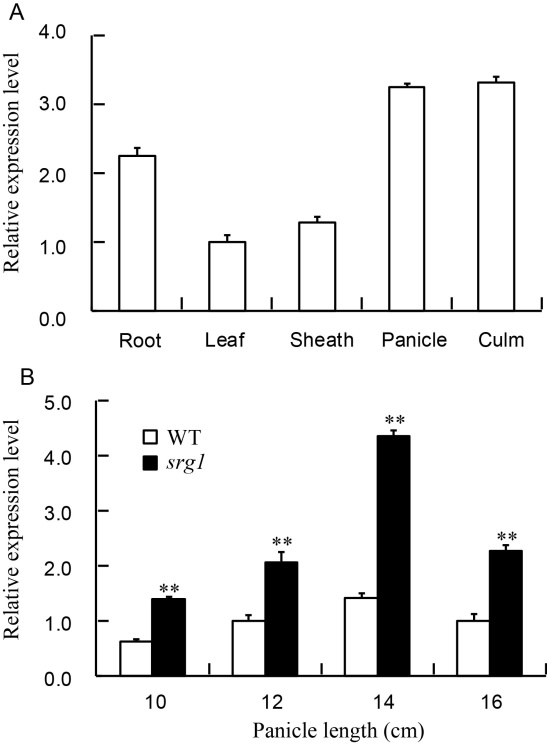
Fig. 5. The expression pattern of SRG1.A, Expression levels of SRG1 in different organs; B, Expression levels of SRG1 in spikelets of different panicle lengths. Rice Actin gene was used as an internal control. Value represents mean ± SD (n = 3); **, P ≤ 0.01.
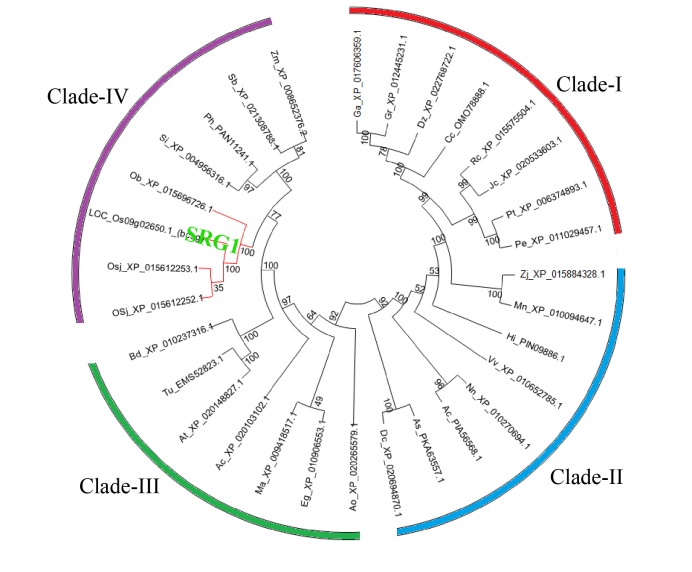
Fig. 6. Phylogenetic tree of homologous proteins of SRG1. Homologs were obtained by a BLASTP in the National Center for Biotechnology Information against the amino acid sequence of SRG1. This analysis was performed by MEGA 6.0 with Neighbor-Joining (NJ) method. The SRG1 is marked with green color and the proteins from Oryza are indicated by red line.
| [1] | Abe Y, Mieda K, Ando T, Kono I, Yano M, Kitano H, Wasaki Y.2010. TheSMALL AND ROUND SEED1 (SRS1/DEP2) gene is involved in the regulation of seed size in rice. Genes Genet Syst, 85(5): 327-339. |
| [2] | Aohara T, Kotake T, Kaneko Y, Takatsuji H, Tsumuraya Y, Kawasaki S.2009. RiceBRITTLE CULM 5(BRITTLE NODE) is involved in secondary cell wall formation in the sclerenchyma tissue of nodes. Plant Cell Physiol, 50(11): 1886-1897. |
| [3] | Cobbe N, Heck M M S.2004. The evolution of SMC proteins: Phylogenetic analysis and structural implications. Mol Biol Evo, 21(2): 332-347. |
| [4] | Duan P G, Rao Y H, Zeng D L, Yang L Y, Xu R, Zhang B L, Dong G J, Qian Q, Li Y H.2014. SMALL GRAIN 1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. Plant J, 77(4): 547-557. |
| [5] | Fan C H, Xing Y Z, Mao H L, Lu T T, Han B, Xu C G, Li X H, Zhang Q F.2006. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet, 112(6): 1164-1171. |
| [6] | Hirano K, Kotake T, Kamihara K, Tsuna K, Aohara T, Kaneko Y, Takatsuji H, Tsumuraya Y, Kawasaki S.2010. Rice BRITTLE CULM 3 (BC3) encodes a classical dynamin OsDRP2B essential for proper secondary cell wall synthesis. Planta, 232(1): 95-108. |
| [7] | Hong Z, Ueguchi-Tanaka M, Shimizu-Sato S, Inukai Y, Fujioka S, Shimada Y, Takatsuto S, Agetsuma M, Yoshida S, Watanabe Y, Uozu S, Kitano H, Ashikari M, Matsuoka M.2002. Loss-of-function of a rice brassinosteroid biosynthetic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem.Plant J, 32: 495-508. |
| [8] | Hong Z, Ueguchi-Tanaka M, Fujioka S, Takatsuto S, Yoshida S, Hasegawa Y, Ashikari M, Kitano H, Matsuoka M.2005. The rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of Arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant Cell, 17(8): 2243-2254. |
| [9] | Hu J, Wang Y X, Fang Y X, Zeng L J, Xu J, Yu H P, Shi Z Y, Pan J J, Zhang D, Kang S J, Zhu L, Dong G J, Guo L B, Zeng D L, Zhang G H, Xie L H, Xiong G S, Li J Y, Qian Q.2015. A rare allele of GS2 enhances grain size and grain yield in rice. Mol Plant, 8(6): 1455-1465. |
| [10] | Huang K, Wang D K, Duan P G, Zhang B L, Xu R, Li N, Li Y H.2017. WIDE AND THICK GRAIN 1, which encodes an otubain-like protease with deubiquitination activity, influences grain size and shape in rice. Plant J, 91: 849-860. |
| [11] | Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B, Onishi A, Miyagawa H, Katoh E.2013. Loss of function of the IAA-glucose hydrolase geneTGW6 enhances rice grain weight and increases yield. Nat Genet, 45(6): 707-712. |
| [12] | Jiang Y H, Bao L, Jeong S Y, Kim S K, Xu C G, Li X H, Zhang Q F.2012. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J, 70(3): 398-408. |
| [13] | Kitagawa K, Kurinami S, Oki K, Abe Y, Ando T, Kono I, Yano M, Kitano H, Iwasaki Y.2010. A novel kinesin 13 protein regulating rice seed length. Plant Cell Physiol, 51(8): 1315-1329. |
| [14] | Kotake T, Aohara T, Hirano K, Sato A, Kaneko Y, Tsumuraya Y, Takatsuji H, Kawasaki S.2011. Rice Brittle culm 6 encodes a dominant-negative form of CesA protein that perturbs cellulose synthesis in secondary cell walls. J Exp Bot, 62(6): 2053-2062. |
| [15] | Li J, Jiang J F, Qian Q, Xu Y Y, Zhang C, Xiao J, Du C, Luo W, Zou G X, Chen M L, Huang Y Q, Feng Y Q, Cheng Z K, Yuan M, Chong K.2011. Mutation of riceBC12/GDD1, which encodes a kinesin-like protein that binds to a GA biosynthesis gene promoter, leads to dwarfism with impaired cell elongation. Plant Cell, 23(2): 628-640. |
| [16] | Li J, Xu Y Y, Chong K.2012. The novel functions of kinesin motor proteins in plants. Protoplasma, 249(2): 95-100. |
| [17] | Li Y H, Qian Q, Zhou Y H, Yan M X, Sun L, Zhang M, Fu Z M, Wang Y H, Han B, Pang X M, Chen M S, Li J Y.2003. BRITTLE CULM1, which encodes a COBRA-like protein, affects the mechanical properties of rice plants. Plant Cell, 15(9): 2020-2031. |
| [18] | Nakagawa H, Tanaka A, Tanabata T, Ohtake M, Fujioka S, Nakamura H, Ichikawa H, Mori M.2012. SHORT GRAIN1 decreases organ elongation and brassinosteroid response in rice. Plant Physiol, 158(3): 1208-1219. |
| [19] | Persson S, Caffall K H, Freshour G, Hilley M T, Bauer S, Poindexter P, Hahn M G, Mohnen D, Somerville C.2007. TheArabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell, 19(1): 237-255. |
| [20] | Preuss M L, Kovar D R, Lee Y R J, Staiger C J, Delmer D P, Liu B.2004. A plant-specific kinesin binds to actin microfilaments and interacts with cortical microtubules in cotton fibers. Plant Physiol, 136(4): 3945-3955. |
| [21] | Reddy V S, Reddy A S N.2002. The calmodulin-binding domain from a plant kinesin functions as a modular domain in conferring Ca2+-calmodulin regulation to animal plus- and minus-end kinesins. J Biol Chem, 277: 48058-48065. |
| [22] | Schmittgen T D, Livak K J.2008. Analyzing real-time PCR data by the comparative C-T method. Nat Prot, 3(6): 1101-1108. |
| [23] | Segami S, Kono I, Ando T, Yano M, Kitano H, Miura K, Iwasaki Y.2012. Small and round seed 5 gene encodes alpha-tubulin regulating seed cell elongation in rice. Rice, 5(1): 4. |
| [24] | Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M.2008. Deletion in a gene associated with grain size increased yields during rice domestication.Nat Genet, 40(8): 1023-1028. |
| [25] | Song X J, Huang W, Shi M, Zhu M Z, Lin H X.2007. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet, 39(5): 623-630. |
| [26] | Tanabe S, Ashikari M, Fujioka S, Takatsuto S, Yoshida S, Yano M, Yoshimura A, Kitano H, Matsuoka M, Fujisawa Y, Kato H, Iwasaki Y.2005. A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant,dwarf11, with reduced seed length. Plant Cell, 17(3): 776-790. |
| [27] | Ueguchi-Tanaka M, Fujisawa Y, Kobayashi M, Ashikari M, Iwasaki Y, Kitano H, Matsuoka M.2000. Rice dwarf mutantd1, which is defective in the alpha subunit of the heterotrimeric G protein, affects gibberellin signal transduction. Proc Natl Acad Sci USA, 97(21): 11638-11643. |
| [28] | van Soest P J, Robertson J B, Lewis B A.1991. Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition.J Dairy Sci, 74(10): 3583-3597. |
| [29] | Vert G, Chory J.2006. Downstream nuclear events in brassinosteroid signalling. Nature, 441: 96-100. |
| [30] | Wang L, Xu Y Y, Ma Q B, Li D, Xu Z H, Chong K.2006. Heterotrimeric G protein alpha subunit is involved in rice brassinosteroid response. Cell Res, 16(12): 916-922. |
| [31] | Wang S K, Wu K, Yuan Q B, Liu X Y, Liu Z B, Lin X Y, Zeng R Z, Zhu H T, Dong G J, Qian Q, Zhang G Q, Fu X D.2012. Control of grain size, shape and quality byOsSPL16 in rice. Nat Genet, 44(8): 950-954. |
| [32] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D.2015. TheOsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat Genet, 47(8): 949-956. |
| [33] | Wang Y X, Xiong G S, Hu J, Jiang L, Yu H, Xu J, Fang Y X, Zeng L J, Xu E B, Xu J, Ye W J, Meng X B, Liu R F, Chen H Q, Jing Y H, Wang Y H, Zhu X D, Li J Y, Qian Q.2015. Copy number variation at theGL7 locus contributes to grain size diversity in rice. Nat Genet, 47(8): 944-948. |
| [34] | Wu Y Z, Fu Y C, Zhao S S, Gu P, Zhu Z F, Sun C Q, Tan L B.2016. CLUSTERED PRIMARY BRANCH 1, a new allele of DWARF11, controls panicle architecture and seed size in rice. Plant Biotech J, 14(1): 377-386. |
| [35] | Xing Y Z, Zhang Q F.2010. Genetic and molecular bases of rice yield. Annu Rev Plant Biol, 61: 421-442. |
| [36] | Xu J D, Zhang Q G, Zhang T, Zhang H Y, Xu P Z, Wang X D, Wu X J.2008. Phenotypic characterization, genetic analysis and gene-mapping for a brittle mutant in rice. J Integr Plant Biol, 50(3): 319-328. |
| [37] | Yamamuro C, Ihara Y, Wu X, Noguchi T, Fujioka S, Takatsuto S, Ashikari M, Kitano H, Matsuoka M.2000. Loss of function of a ricebrassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell, 12(9): 1591-1605. |
| [38] | Yu H P, Ren D Y, Zhu Y Z, Xu J M, Wang Y X, Liu R F, Fang Y X, Shi Z Y, Pan J J, Lu M, Ma B J, Hu J, Rao Y C.2016. Multi-tillering dwarf1, a new allele of brittle culm 12, affects plant height and tiller in rice. Sci Bull, 61(23): 1810-1817. |
| [39] | Zhang B C, Liu X L, Qian Q, Liu L F, Dong G J, Xiong G Y, Zeng D L, Zhou Y H.2011. Golgi nucleotide sugar transporter modulates cell wall biosynthesis and plant growth in rice.Proc Natl Acad Sci USA, 108(12): 5110-5115. |
| [40] | Zhang M, Zhang B C, Qian Q, Yu Y C, Li R, Zhang J W, Liu X L, Zeng D L, Li J Y, Zhou Y H.2010. Brittle Culm 12, a dual-targeting kinesin-4 protein, controls cell-cycle progression and wall properties in rice. Plant J, 63(2): 312-328. |
| [41] | Zhang Q, Shen B Z, Dai X K, Mei M H, Saghai Maroof M A, Li Z B.1994. Using bulked extremes and recessive class to map genes for photoperiod-sensitive genic male sterility in rice.Proc Natl Acad Sci USA, 91(18): 8675-8679. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||